Difference between revisions of "SUIT-024 O2 ce-pce D056"
Moelk Sophia (talk | contribs) |
Beno Marija (talk | contribs) |
||
| (2 intermediate revisions by 2 users not shown) | |||
| Line 5: | Line 5: | ||
|application=O2 | |application=O2 | ||
|SUIT number=D056_ce1;1Dig;1PM;2T;2D;3Omy;4Ama | |SUIT number=D056_ce1;1Dig;1PM;2T;2D;3Omy;4Ama | ||
}} | }} | ||
::: '''[[Categories of SUIT protocols |SUIT-category]]:''' N(PM) | ::: '''[[Categories of SUIT protocols |SUIT-category]]:''' N(PM) | ||
::: '''[[SUIT protocol pattern]]:''' Bidirectional linear ce1;1Dig;1PM;2T;2D;3Omy;4Ama | ::: '''[[SUIT protocol pattern]]:''' Bidirectional linear ce1;1Dig;1PM;2T;2D;3Omy;4Ama | ||
| Line 24: | Line 23: | ||
:::* This protocol allows to analyse the presence of ATPases in [[mitochondrial preparations]] and is useful as a quality control of the sample preparation. | :::* This protocol allows to analyse the presence of ATPases in [[mitochondrial preparations]] and is useful as a quality control of the sample preparation. | ||
:::+ Different combinations of substrates and inhibitors can be used depending on the specific aims, ''e.g.'': PGM, PM, GM, MnaPM or other combinations for N-pathway control state; S or RotS for S-pathway control state. | :::+ Different combinations of substrates and inhibitors can be used depending on the specific aims, ''e.g.'': PGM, PM, GM, MnaPM or other combinations for N-pathway control state; S or RotS for S-pathway control state. | ||
:::+ [[LEAK]] | :::+ [[LEAK respiration]] overestimation is prevented in the presence of [[Oligomycin]]. | ||
:::+ Reasonable duration of the experiment. | :::+ Reasonable duration of the experiment. | ||
:::- This protocol does not include internal [[ | :::- This protocol does not include internal [[Electron transfer pathway]] control steps. | ||
:::- CIV activity is not measured, to save experimental time. | :::- CIV activity is not measured, to save experimental time. | ||
:::- Careful washing is required after the experiment to avoid carry-over of oligomycin. | :::- Careful washing is required after the experiment to avoid carry-over of oligomycin. | ||
== Compare SUIT protocols == | == Compare SUIT protocols == | ||
== Chemicals and syringes == | |||
{{Template:Chemicals ce}} | |||
{{Template:Chemicals SUIT-024}} | |||
::: Suggested stock concentrations are shown in the specific DL-Protocol. | |||
== References == | == References == | ||
Latest revision as of 12:36, 25 November 2020
Description
Abbreviation: N(PM)
Reference: Determination of the presence of ATPases in mitochondrial preparations - SUIT-024
SUIT number: D056_ce1;1Dig;1PM;2T;2D;3Omy;4Ama
O2k-Application: O2
- SUIT-category: N(PM)
- SUIT protocol pattern: Bidirectional linear ce1;1Dig;1PM;2T;2D;3Omy;4Ama
SUIT-024 O2 ce-pce D056 is a protocol for permeabilized cells, designed to quantify the ATPase activity in the sample. When isolating mitochondria, contamination with other membranous organelles containing ATPases (e.g., plasma membrane and endoplasmic reticulum) can occur. In experiments with tissue homogenates and permeabilized cells, it is expected to have active ATPases in the respiration medium. When ATPases are active and residual endogenous adenylates are present in the sample, ATP can be consumed and converted to ADP, which will stimulate respiration. This may lead to an overestimation of LEAK respiration - L(n). Therefore, if the samples contains ATPases/adenylates, it might be necessary to assess LEAK respiration in the presence of ATPases inhibitors or oligomycin - L(Omy). This protocol can be used as a quality control of the mitochondrial preparation.
In the DatLab software, SUIT-024 DLP file is provided for ce-pce and the category N(PM). For using this protocol with other sample preparation or substrate/inhibitor combinations, a personalized DLPU can be created.
Communicated by Garcia-Souza LF (last update 2019-05-14)
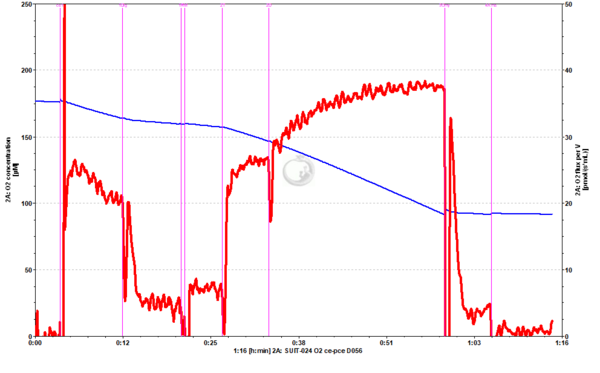
Representative traces
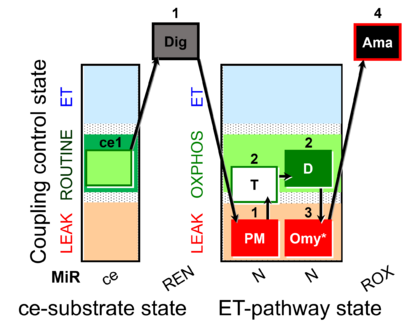
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| ce1 | ROUTINE | ce1
| ||
| 1Dig | REN | ce1;1Dig
|
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1PM | PML(n) | N | CI | 1PM
|
| 2T | PML or PMP | N | CI | 1PM;2T
|
| 2D | PMP | N | CI | 1PM;2T;2D
|
| 3Omy | PML(Omy) | 1PM;2T;2D;3Omy
| ||
| 4Ama | ROX | 1PM;2T;2D;3Omy;4Ama
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- This protocol allows to analyse the presence of ATPases in mitochondrial preparations and is useful as a quality control of the sample preparation.
- + Different combinations of substrates and inhibitors can be used depending on the specific aims, e.g.: PGM, PM, GM, MnaPM or other combinations for N-pathway control state; S or RotS for S-pathway control state.
- + LEAK respiration overestimation is prevented in the presence of Oligomycin.
- + Reasonable duration of the experiment.
- - This protocol does not include internal Electron transfer pathway control steps.
- - CIV activity is not measured, to save experimental time.
- - Careful washing is required after the experiment to avoid carry-over of oligomycin.
Compare SUIT protocols
Chemicals and syringes
| Step | Chemical(s) and link(s) | Comments |
|---|---|---|
| 1Dig | Digitonin (Dig) |
| Step | Chemical(s) and link(s) | Comments |
|---|---|---|
| 1PM | Pyruvate (P) and Malate (M) | |
| 2T | ATP (T) | |
| 2D | ADP (D) | |
| 3Omy | Oligomycin (Omy) | |
| 4Ama | Antimycin A (Ama) |
- Suggested stock concentrations are shown in the specific DL-Protocol.
References
MitoPedia concepts: SUIT protocol, SUIT A, Find
MitoPedia methods:
Respirometry