- high-resolution terminology - matching measurements at high-resolution
User talk:Sigl Reinhard/My sandbox flux analysis
General
DatLab sets a novel standard in high-resolution respirometry, for on-line analysis of oxygen flux measured in the Oroboros O2k, and of other signals obtained in the O2k-MultiSensor System. Oxygen flux can be plotted with or without O2 kbackground correction, is instantaneously expressed per mass of sample or per number of cells. Various sections on the plot of oxygen flux are marked, and corresponding average values are viewed in a table which can be simply exported into Excel or SigmaPlot. Instrumental and experimental parameters are summarized in a protocol, which can be printed or saved as a pdf file. These new features provide the basis for combining high-resolution with instant and user-friendly analysis. All analyses can be performed real-time or disconnected from the O2k.
- » Product: DatLab, Oroboros O2k, O2k-Catalogue
DatLab 7
Online manual:
- 1. Overview:
- A demonstration experiment, performed during an O2k-Workshop on high-resolution respirometry, is used as an example for application of DatLab and DatLab-Excel templates MiPNet08.09. All analyses can be performed real-time or disconnected from the O2k.
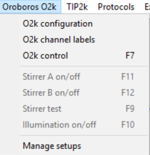
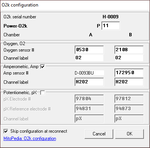
- O2k Click on 'Oroboros O2k' and select 'O2k configuration'. In the O2k-window which appears select the channels of interest.
- O2k-O2 applications: Deselect all O2k-MultiSensor channels.
- O2k-MultiSensor applications: Select O2 channel and the specifically O2k-MultiSensor channels (Amp or pX).
- 2. Oxygen flux of a biological sample
- 2.1 O2k-Demo file
-
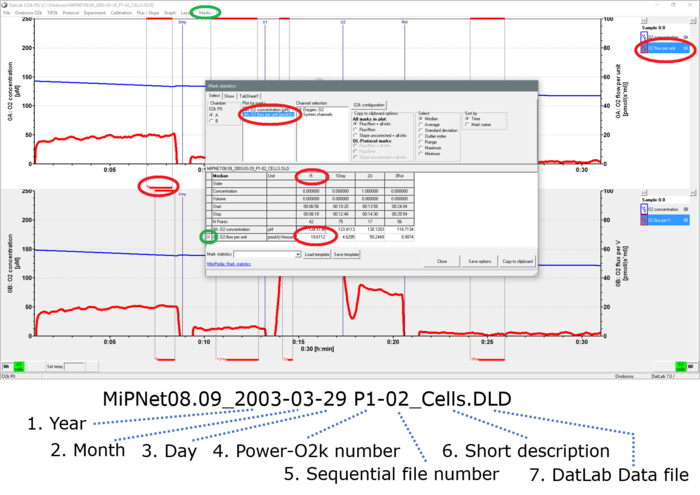
- MiPNet08.09_IntactCells.DLD O2 concentration (blue, Y1 axis) and O2 flux (red, Y2 axis) as a function of time. Data recording was started (Connect [F7]) after adding a cell suspension at a density of 1∙106 cells/mL. Events are shown by vertical lines, with the <Event name> below. Marks are shown by horizontal bars between two vertical lines, with the Mark name in the lower bar. To be conplete, here is the according DatLab-Analysis template: MiPNet08.09_O2k-Analysis_Cells.xlsx.
- Edit experiment [F3] Select the Sample Unit Million cells. Enter the cell density [1.000]. The amount of cells in the chamber is then shown below, depending on the chamber volume (2.00 mL).
- Calibration [F5] See MiPNet19.18D O2k-calibration. The calibration from the previously saved file is available as a default. The oxygen solubility factor, FM, is 0.89 for culture medium (RPMI).
- View protocol Select Experiment and then Experimental Log or press [Ctrl+F3] and Preview. The protocol has been saved as a pdf file “MiPNet08.09_IntactCells.pdf”.
- Graph layout 04a Flux per volume. This layout provides a plot of volume-specific respiratory oxygen flux, which is most relevant to evaluate experimental details, for instance the flux measured in relation to the sensitivity of the instrument (1 pmol∙s−1∙cm−3; [MiPNet18.10]). This plot is also chosen, when measurements of sample concentration are available at a later stage only (in DatLab, press [F6] and Info for further information).
- 2.2 Flux per mass or flow per cell
- Expressions of oxygen flux (corrected for O2-background)
Volume-specific flux JO2
[pmol∙s-1∙ml-1]: The experimental flux per unit of chamber volume is the basis for expressing respiration in a variety of units. Flow IO2
[pmol∙s-1∙10-6 cells]: A system-specific quantity, in contrast to the size-specific quantities. Mass-specific flux JO2
[pmol∙s-1∙mg-1] Flux control ratio FCR
Normalized flux, dimensionless, relative rate.
- Graph layout 05a Specific flux is used for plotting respiration per unit sample (Y2 axis), in units defined in “Experiment/Edit” [F3] (oxygen flow per million cells, flux per biomass or protein [mg/mL]). Respiratory flux per chamber volume is converted to an extensive quantity (flow; per cell) or a size-specific quantity (flux; per mg cell protein or mass). Flow and flux are always corrected for instrumental O2 background, using the parameters entered in the “O2 slope” tab of “Flux/Slope”. Press [F6] and Info for information.
- 3. O2-instrumental background oxygen flux
- 4. References
PDF manual:
MitoPedia O2k and high-resolution respirometry:
O2k-Open Support
- DatLab plots of O2 flux per mass or flow per cell (per unit sample) remain shown per volume
- Problem: After selecting a plot of the O2 flux per mass or flow per cell (per unit sample or the), the plot in the graph remains shown as flux per volume. Neither selecting the plot manually in 'Graph - Select Plots' (in the menu) nor choosing the appropriate layout from the menu 'Layout' helps.
- Cause
- Sample Concentration\Amount inside the 'Edit Experiment' window (menu 'Experiment - Edit') equal to 0
- The information of the Sample Unit\Concentration\Amount inside the 'Edit Experiment' window (menu 'Experiment - Edit') hasn't been saved yet. Although the values inside this window are taken from the last experiment (measured with this DatLab installation, where the 'Save' button inside the 'Edit Experiment' has been pressed) and displayed at the beginning of a next experiments, they are not automatically saved. - This is only relevant up to DatLab version 6.
- Solution
- Enter the Concentration\Amount different from 0
- Inside the 'Edit Experiment' window (menu 'Experiment - Edit') review (or enter) the sample information (Unit\Concentration\Amount), click on the 'Save' button - then the appropriate plot can be selected (if the Concentration\Amount is not equal to 0).
- ~ Capek Ondrej 15:24, 2 April 2015 (CEST)
- Fluxes are different in left and right chamber
- Question: In our experiments we get consistently higher fluxes in chamber A as compared to chamber B.
- Solutions
- To exclude instrumental causes please follow the procedure described in the section Calibration and quality control of the OroboPOS (O2k-SOP) in MiPNet06.03 POS-calibration-SOP.
- Possible instrumental and protocol related causes for fluxes differing between chambers:
- Wrong O2 calibration in one chamber, see MiPNet06.03 POS-calibration-SOP.
- Wrong chamber volume calibration: The total amount of oxygen in the chamber with the smaller volume will be smaller. Therefore, the O2 concentration will drop faster resulting in a higher flux, see MiPNet19.18A O2k-start.
- Wrong background parameters causeing an O2 concentration-dependent offset to the flux, the value of the offset will be independent of the absolute value of the flux. E.g. a constant offset of 10 pmol / s ml at ca 100 µM O2 concentration and 20 pmol/s ml at 300 µM O2 concentration could be caused by wrong background parameters, but a difference of always e.g. 20% at very different absolute values of the flux can not be caused by wrong background parameters. Check the background parameters (perform an instrumental O2 background experiment). See MiPNet14.06 Instrumental O2 background.
- Sample injection: One filling of a syringe is used to inject the sample into several chambers. Sedimentation starts to occur immediately and more sample will be injected into the first chamber than in any subsequent chambers. A remedy we apply is to fill the syringe with sample sufficient for 3 chambers and inject the first aliquot back to the sample stock solution before quickly injecting the rest into the chambers. Independently of this we recommend to randomize chamber assignment to prevent systematic errors.
- Hydrophobic inhibitors, see MiPNet19.03 O2k-cleaning and ISS.
- Biological contamination, see Biological contamination.
- Hardware defects: While hardware defects (sensors, electronics,..) can obviously have many negative effects (noise signal, slow response, no signal,..) it is difficult to see how a real hardware defect can cause a systematic error in flux calculation once a correct calibration of the O2 sensor (at air saturation and at zero oxygen) was obtained.
Labels: MiParea: Respiration, Instruments;methods
HRR: Oxygraph-2k, O2k-Manual, O2k-Protocol
O2k-Core, DatLab, DL6, DL6a7, SeriesG